
A total of 150 isolates were considered significant by the criteria mentioned, which accounts for about 79% of the nonfermenters causing infection. The remaining 91 specimens (48.2%) showed monomicrobial infection. coli and S.aureus were commonly associated. Ninety-eight specimens (51.8%) showed polymicrobial infection where nonfermenters were isolated along with other organisms, of which E. A total of 193 NFGNB were isolated from 189 clinical specimens as four samples yielded two NFGNB. NFGNB were isolated from 189 out of 4248 clinical specimens accounting for an isolation rate of 4.5%. aeruginosa ATCC 27853 were used as the control strains. The results were interpreted as per the Clinical and Laboratory Standards Institute (CLSI) guidelines.Į. The different antimicrobials tested were imipenem, piperacillin, ticarcillin, amikacin, ciprofloxacin, ceftazidime, cefepime, cefoperazone, ceftriaxone and cotrimoxazole. The sensitivity test was performed with the help of the Kirby-Bauer disc diffusion method using commercially available discs (Hi-media). The clinical criteria included the presence of risk factors, such as, bronchiectasis, cystic fibrosis, underlying pneumonia, any malignancy, on mechanical ventilation, and other immunosuppressive conditions. The laboratory criteria included: The presence of pus cells along with gram-negative bacilli in the stained smear from the sample, monomicrobial infection, isolation of the same organism from a repeat sample, leukocytosis, and relevant radiological evidence (in cases of VAP). The clinical significance of the isolated NFGNB was assessed retrospectively by analyzing the case sheets for a combination of relevant laboratory and clinical criteria. pseudomallei is oxidase +ve, OF: Oxidation-fermentation 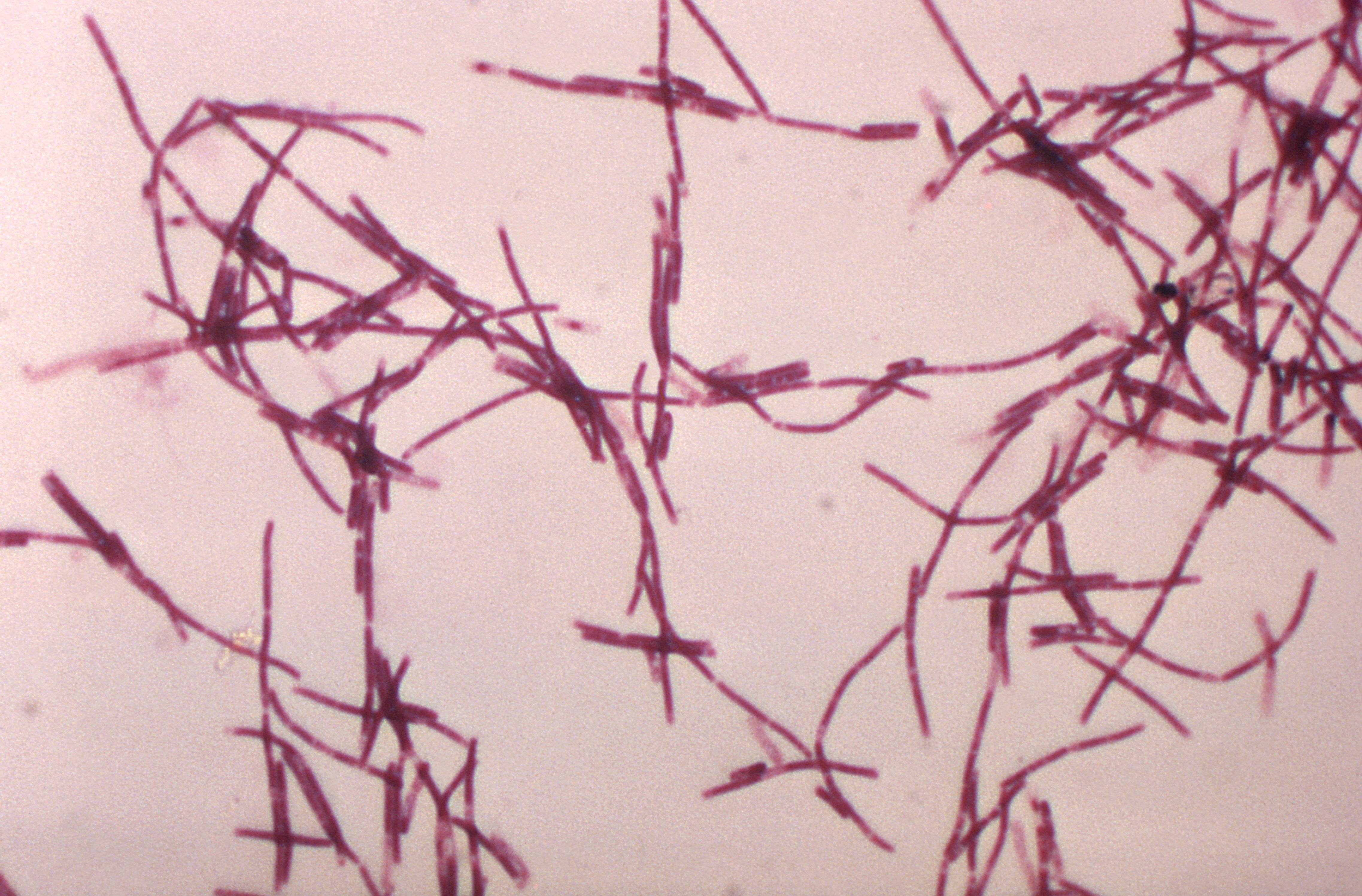
Scheme used in the study for identification of nonfermenting gram-negative bacilli * Burkholderia species vary in their oxidase reaction: Burkholderia gladioli is oxidase -ve, B. The scheme for identification of NFGNB is shown in Figure 1. The characters assessed included morphology on Gram's stain, motility, pigment production, oxidase production, OF test (Hugh-Leifson's medium) for glucose, lactose, sucrose, maltose, mannitol, and xylose, growth on 10% lactose agar, lysine decarboxylase test, and gelatin liquefaction test. All the organisms that grew on Triple Sugar Iron agar and produced an alkaline reaction were provisionally considered to be NFGNB and identified further by using a standard protocol for identification. The organisms isolated were identified using the appropriate biochemical tests. These samples were plated on blood agar, chocolate agar, and Mac Conkey's agar, and incubated at 37☌ for 18-24 hours. The study was also done to assess their clinical significance and antimicrobial susceptibility pattern, and to identify the various healthcare-related infection they cause.Ī total of 4248 clinical specimens were received in the laboratory during November 2005 and July 2006, which included 1428 urine, 1124 pus, 1035 blood cultures, collected in a brain-heart infusion broth, 365 sputum and respiratory samples, and 296 body fluids. Hence, this study was undertaken to identify the various nonfermenters isolated from patients admitted to our hospital, a tertiary care center, at Kolar. There are very few studies from India wherein the various NFGNB, isolated from patients’ samples, have been identified and their clinical significance assessed. NFGNB are innately resistant to many antibiotics and are known to produce extended spectrum ß-lactamases and metallo ß-lactamases. They have been incriminated in infections, such as, septicemia, meningitis, pneumonia, urinary tract infections (UTI), and surgical site infections (SSI). In recent years, due to the liberal and empirical use of antibiotics, NFGNB have emerged as important healthcare-associated pathogens. NFGNB are known to account for about 15% of all bacterial isolates from a clinical microbiology laboratory. They occur as saprophytes in the environment and some are also found as commensals in the human gut. Nonfermenting gram-negative bacilli (NFGNB) are a taxonomically diverse group of aerobic, nonsporing, bacilli that either do not utilize glucose as a source of energy or utilize it oxidatively.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
